
We are innovating in the protein design field. Our business activity has started in November 2024!
By combining physics, robotics-inspired algorithms, deep/machine learning and automated reasoning, we design improved proteins from first principles and provide customers with optimized molecules for all type of applications.

Our pure in silico design processes avoid most of the expensive and time consuming experimental essays. Only carefully selected designs need to be tested.

Our AI-powered design method is able to confer new, desirable properties, providing sequences that evolution has not been able to explore.

With its extreme flexibility, our design engine can offer customized solutions that match your needs: specificity, stability, sequence composition, activity,...

Every design problem is different. We offer a variety of services, adapted to a range of different requirements, from the simplest to the most challenging ones.
Physics, Machine Learning and Robotics united by Automated reasoning.
Our technology relies on a disruptive combination of physics-based and robotics-inspired modeling; machine and deep learned information extracted from Nature and experimental results, all united by a dedicated automated reasoning engine that efficiently and optimally integrates these different sources of information with custom-defined sequence requirements.
Thanks to this unique combination of features, various protein properties can be targeted for optimization, including thermo-resistance, activity, solubility, specificity, selectivy or affinity.
Starting from fundamental principles and universal data, our technology applies on orphan, de novo designed, as well as existing proteins, including enzymes, binders or self-assembling systems
It can contribute to improve protein-based processes in various fields, including Biotechnology, Chemistry, Bioenergy, Health, Environment and Food.

When structural information is available, we rely on the most recent atomic and quantum molecular modeling force-fields, and scoring functions to choose an amino acid sequences that will make your protein real, effective and resistant to the various stresses it may have to sustain during its existence.
Using all the data available in structure and sequence databases, we exploit the most recent generative Machine and Deep learning technology to extract information on sequence, structure and function relationships from Nature and other experimental data. Thanks to this, ill-defined properties such as expressability can also be optimized.
Proteins can be modeled as complex poly-articulated robotic systems with many degrees-of-freedom. We leverage recent and fast robotics-inspired algorithms to efficiently capture the crucial flexibilities of target proteins, from specific loop movements to essential ligand access/exits paths.
Our final sequence design engine relies on state-of-the-art AI automated reasoning technology to integrate all the information provided by Physics, Machine learning and Robotics with customer specific requirements to efficiently produce a library of diverse sequences that can also account for multiple targeted conformational states.
A team of dedicated experienced professionals.

Co-founder
Sophie is a Research Director at INRAE, specialized in protein modeling and design. She has a long experience of interactions with academic labs and companies for the tailor-made conception of proteins with improved and new capabilities for Biotechnology and Health. Her expertise covers computational protein design, methodological developments for structural bioinformatics, structure-function relationship studies, and rational protein engineering. She has co-authored over 60 scientific papers and 5 patents and has contributed to the development of several molecular and design methods (AI-based).

Co-founder
Juan is a Research Director at CNRS and a computer scientist specialized in algorithmic robotics. He is one of the pioneers in the development of structural bioinformatics approaches based on algorithms originating from robotics and artificial intelligence. During the last 15 years, he has conducted fruitful interdisciplinary research in this area in collaboration with bio-chemists, biologists and physicists. He has co-authored over 90 scientific papers and coordinated the development of several software packages and tools, in particular MoMA.

Co-founder
Thomas is a Research director at INRAE, a computer scientist specialized in hybrid systems combining logical and numerical automated reasoning with probabilistic machine learning and deep learning methods in AI. He also contributed to their extension and applications in computational biology from genetic, genomics to structural biology. He has co-authored more than 160 scientific papers and several original bioinformatics and core-AI software. He is a Fellow of the Association for the Advancement of AI and of the European Association for AI.
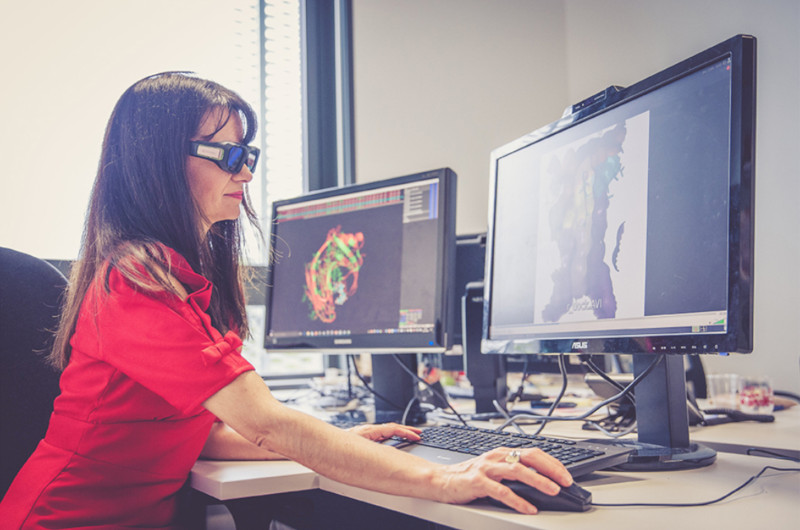

We are improving our technology and its scope of application every day. Contact us, we may well already have the capacity to build your solution.
Send us a message, we will call back later
This site is protected by reCAPTCHA and the Google Privacy Policy and Terms of Service apply.